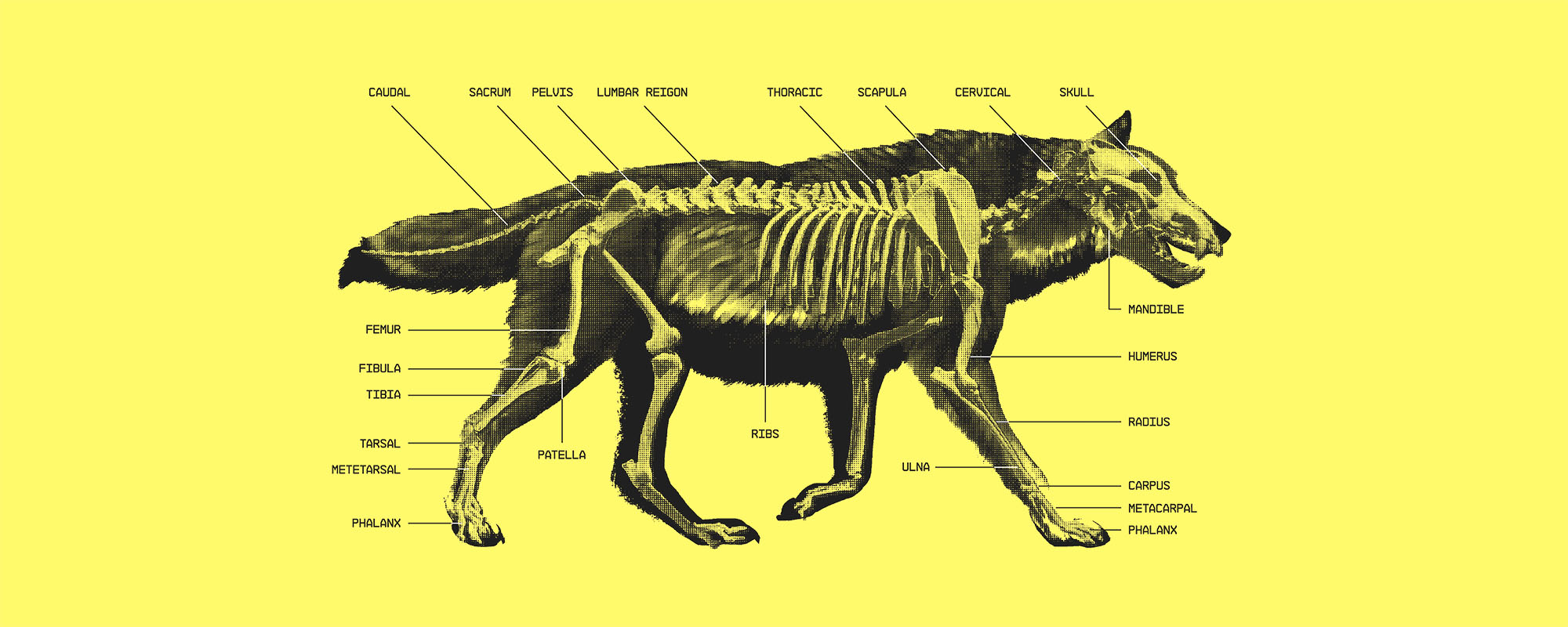
De-extinction: The Case of the Dire Wolf? – The ‘de-extinction’ of the dire wolf has been one of the highest-profile scientific achievements of 2025. But the impression cultivated online is misleading – the dire wolf has not returned; it is being imitated. Five professors from the University of Malta have reacted to this complex story. In Part IV, Prof. Louis-F. Cassar considers the moral questions raised by a de-extincted future.
Continue readingCutting EDGE Research: the Genomes of Maltese Plants
Conservation is a 21st century hot topic. It is a top priority worldwide to ensure the lasting protection of the planet and its natural resources. But how can we even begin to conserve our natural environment if we don’t understand it? Ines Ventura investigates.
Continue readingDNA spacewalk
Can bacteria grow in space? How do cells respond to radiation? Geneticist Prof. Joseph Borg realised that to find out, he must make some friends outside his discipline.
Continue readingKeep Calm and Factor This
Author: Malcolm Pace
People suffering from haemophilia, an inherited disorder, will bleed for much longer after an injury, bruise easily, and risk internal bleeding in joints or brain. This is a widely studied condition with unique variations in the disorder to Malta. While pursuing an M.Sc. in Applied Biomedical Science, I have profiled and found the contributing genes in all Maltese patients and compared them to patients abroad.
Continue readingLiving with a rare disease
Author: Clayton Axiak
Picture yourself waking up one morning with a severe, relentless itch that no clinician or diagnostic tool can understand. Your life would be thrown off kilter. Quality of life would suffer financially, psychologically, and socially as you try to look for a glimmer of light at the end of the tunnel. This is what life is like for most people living with a rare disease.
Often barraged with terms like ‘unknown’ or ‘undiagnosed’, matters can get even more challenging when the condition becomes more elusive or develops life-threatening consequences. And all of this is exacerbated by inequities in treatment and high costs of the few existing drugs that are available.
By EU standards, a rare disease is one that affects fewer than one in 2,000 individuals. And these ‘less common’ ailments are difficult to raise monies for to research, leaving large gaps in scientific and medical literature. One such disease is the poorly understood Idiopathic Hypogonadotropic Hypogonadism (IHH).
Characterised by the absence of puberty and infertility, IHH can be compounded by potentially severe characteristics such as congenital heart disease, osteoporosis at a young age, and early onset of Alzheimer’s disease.
Its cause is usually a genetic anomaly, but a single genetic change can affect two people very differently. This gives rise to an unparalleled complexity that makes the cause harder to decipher. Symptoms are not clear-cut and sometimes mask the actual underlying cause, bringing about misdiagnosis and delayed treatment. Timely diagnosis is crucial for successful treatment that enables the patient to achieve puberty and induce fertility. But this is not always possible.
Under the guidance of Dr Rosienne Farrugia, I am currently analysing and expanding upon a preliminary assessment of IHH in Malta using high-throughput sequencing (HTS) technology (conducted by Adrian Pleven). With HTS, we can read a person’s entire DNA sequence and attempt to identify differences in the DNA code which lead to such diseases.
What the team has found is that some genetic variants typical of IHH are more common in the Maltese population when compared to mainland Europe and African populations. This is likely due to the reduced genetic variation of our population, shaped by successive events of population reduction and expansion throughout our history.
By mapping the genetic cause of diseases prevalent on our islands, we can help medical consultants to employ specific screening tests that are tailored for local patients suffering from IHH. Such advancements in genomic technology and personalised medicine can make a huge impact on people’s lives. And not only to those suffering from IHH; researching one disease, however rare it may be, can shed light on mechanisms that prove useful in treating many others, ensuring that when it comes to health, no one is left behind.
This research project is being carried out as part of a Ph.D. program in Applied Biomedical Sciences at the Faculty of Health Sciences.
Analysing the building blocks
Gene sequencing for disease treatment
A staggering amount of diseases can be traced back to a genetic cause. Dr Rosienne Farrugia talks to THINK about her team’s efforts to use genome sequencing to eventually secure timely treatment for some very serious conditions.
Continue readingAspirin and Cancer
Aspirin is often considered a wonder drug due to its versatile use in treating inflammation, reducing pain, and helping to prevent heart-related disease. However, there is more to it. Aspirin is actually cancer-preventive. Studies have shown that a daily low dose of aspirin, medically prescribed for more than five years, lowers the risk of cancer-related deaths by at least 30%. So, should we all start taking aspirin on a daily basis to lower our chances of getting cancer?
No, not exactly. This is because many aspects of aspirin’s cancer-preventive effects are still poorly understood. Particularly, researchers have not yet pinpointed what enables aspirin to selectively kill early-stage cancer cells and not healthy cells. This is the scope of the research currently being carried out at the Yeast Molecular Biology and Biotechnology Laboratory (headed by Prof. Rena Balzan).

The secret behind aspirin’s tendency to kill certain cells but not others seems to lie in the physiology of the exposed cells. Aspirin exploits the natural differences between healthy and cancerous cells to eliminate malignant cells before they can take over.
Oxygen, if transformed into ‘Reactive Oxygen Species’, is known to cause DNA mutations that can lead to cancer. Through this research, we studied mutated yeast cells which are a relevant model of early-stage cancer cells due to their low tolerance to oxygen-associated stress. We then identified genes in these mutant yeasts which are affected by aspirin.
One of aspirin’s targets is a key metabolite required for the production of energy-rich compounds vital for cell survival. We found that aspirin creates a shortage of this metabolite in mutated yeast cells, causing them to run out of energy and die.
This implies that early-stage human cancer cells may suffer a similar fate and, more importantly, partly explains how aspirin prevents tumour formation. Such knowledge may prove useful in the development of novel anti-cancer treatments.
This research was carried out as part of Project “R&I-2015-001”, financed by the Malta Council for Science & Technology through the R&I Technology Development Programme. This research is being carried out as part of Azzopardi’s Ph.D. project at the Centre for Molecular Medicine & Biobanking and the Department of Physiology & Biochemistry, University of Malta
Author: Maria Azzopardi
Luminex xMAP®: Enhanced lab efficiency
Stereotypical depictions of researchers involve crazy hair, oversized goggles, shabby lab coats, and loads of test tubes. While the first three may be exaggerated, the sheer volume of tubes and wells needed in a lab cannot be overstated, especially when the lab is dedicated to anything biological.
One tissue sample can be used for a gamut of tests, all of them attempting to identify something different in it, be they antibodies, DNA, or RNA (biomarkers). Often, many samples are required due to all the tests needed to highlight the variations in those biomarkers. But the size of samples is now decreasing thanks to machines like the Luminex System running xMAP technology.
The Luminex System is a research/clinical diagnostics platform that allows detection of multiple analytes in a single well of a microtiter plate—100 or more reactions using a single drop of fluid.
Multiplex assays are widely used in experiments investigating the characteristics of molecules within a biological sample. This approach can be used to see whether an experimental treatment works, or what changes a DNA mutation causes in the molecules or molecular pathways within cells.
In real terms, this machine allows for analyses to be done to determine whether or not a patient has a particular disease or gene variant in their blood that would prevent a drug from being effective. It also allows them to determine the ideal dosage for those drugs. The machine can also be used to identify and characterise viral infections.
A particular research group at the University of Malta, headed by Prof. Godfrey Grech, has used Luminex xMAP technology to develop novel markers which are allowing them to classify a subset of triple-negative breast cancer
patients.
By identifying these biomarkers, it may be possible in future to detect the disease earlier and give patients better-targeted therapy.

Author: Prof. Godfrey Grech
Shifting power dynamics in genomic research
Donation of human tissues has been at the core of medical research in Europe since religious restraints were relaxed towards the end of the Middle Ages, making this altruistic act socially and morally acceptable. Altruism is the key word—the individual donates their tissues for the greater good: for the advancement of science, the extension of boundaries of medical knowledge, and for the potential to deliver better medical care. The individual donating their tissues or blood does so with no expectation of return or personal gain, and hands over sample ownership to the scientific community in confidence that ethical use will be made of it.
Many would assume this is still the case… but is it? Genomic research has opened up new and exciting avenues for medical research that have the potential to dramatically transform the way medical practitioners strategically select treatment regimes. Personalised medicine is no longer a pipe dream, it is clearly in the cards—the pack has been dealt, and the game is well underway.
The game-changer here is the genome. The established practice of donors passively handing over their samples of blood or tissues for research is impacted by the fact that DNA is in focus here. Their DNA.
Should this make any difference? Some argue that the fundamental process of donating for the greater good, with no consequent active involvement, is still the cornerstone of the process. There are, however, important shifts in the current social and cultural context that impact the researcher-donor dynamic. Two of these are of particular relevance here. The first is knowledge: the accessibility (via the web) of information about the research process, its impacts and applications, have led to the burgeoning new field of citizen engagement in research. Genomic research, in particular, attracts participants who are personally motivated in the process, with research into genetically linked diseases offering hope for their relatives and future generations. The second shift is that of rights over personal data. Individuals’ rights to the control of their personal data are to be entrenched in the General Data Protection Regulation in May 2018. This EU regulation will sanction a process that is already clear in practice: the trend for individuals to demand control over the ways their personal data are used and stored.
This is where the detached and passive sample ‘donor’ exits stage left, and the research participant takes the limelight. Cynics might claim that nothing much has changed. Samples are collected with consent and research continues, but there is an exciting difference in the dynamics involved. The participant now has the potential to remain engaged by using a process of web-based consent. The various versions of this eConsent or ‘dynamic consent’ that are now available offer the participant and researcher a real-time channel of communication which enables optimum use and reuse of the sample and the accompanying personal data that make it so valuable. The biomedical samples now come with persons digitally-attached: individuals who actively engage in the research process by offering flexible consent via web-based tools. When pooled and stored within biobanks, these samples offer a viable source of DNA with real potential for wide use in research.
Within this scenario, ethicality is enhanced as banked anonymised samples can be used to their full potential and shared, with consent, within the research community. In this way doing justice to the individual participant’s key motivation: to advance scientific knowledge for the benefit of future generations.
Author: Dr Gillian M. Martin
Special Feature – The Maltese Genome
DNA is what life is made of. Found in every cell of the human body, it has sent criminals to jail and been the focus of controversial court cases. Dr Jean Buttigieg discusses these legal and ethical issues. DNA has also transformed the meaning of being human, with traits from disease to intelligence all linked to it. DNA is changing the world.